FUMA GWAS
Functional Mapping and Annotation of Genome-Wide Association Studies
Announcements
FUMA has been updated to version 2.0.0.
Please navigate to the wiki page for an overview of the updates.
IMPORTANT: Please make sure to read the documentation carefully before submiting your jobs. If you need assistance, please post your questions on FUMA GWAS users with your job type (snp2gene, gene2func, celltype, flames, or xqtls) and your jobID.
As this is a major update, expect that FUMA will be down periodically for bug fixes. Please be sure to always download your files after the analysis is done.
1. We will perform an update and maintenance on FUMA starting from Sunday May 17 2026. During the maintenance, FUMA is not accessible. Expect up to 2 weeks of down time. Please make sure to download files needed for your analyses from FUMA before this date. Any QUEUED jobs would be stopped before the maintenance.
2. Any SNP2GENE jobs that were created prior to Jan 01 2023 will be removed from the FUMA server during the maintenance. This does not apply to public jobs.
About FUMA
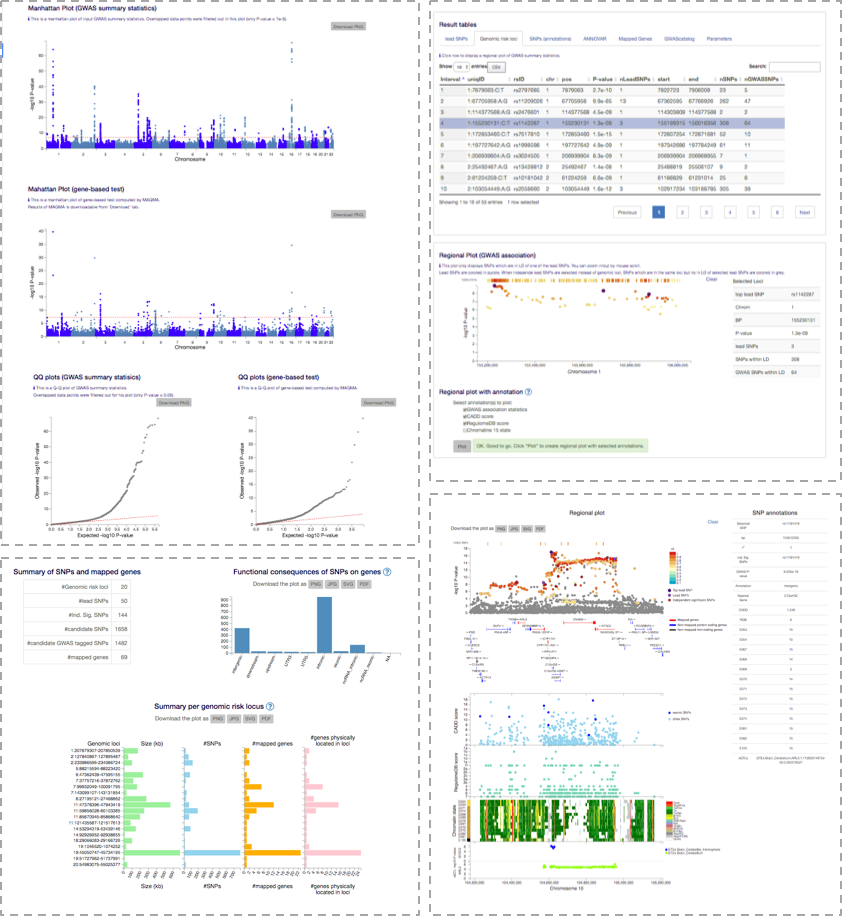
FUMA is a platform that can be used to annotate, prioritize, visualize and interpret GWAS results.
The SNP2GENE module takes GWAS summary statistics as an input,
and provides extensive functional annotation for all SNPs in genomic areas identified by lead SNPs.
The GENE2FUNC module takes a list of gene IDs (as identified by SNP2GENE or as provided manually)
and annotates genes in biological contexts.
The Cell Type module takes MAGMA gene analysis result (as an output from SNP2GENE or as provided manually) and predicts relevant cell types.
The FLAMES module identifies effector genes per genomic risk locus defined from SNP2GENE outputs.
The QTLs Analysis module investigates the potential functional mechanisms underlying GWAS associations by integrating with various QTLs.
To submit your own GWAS, login is required for security reason.
If you have not registered yet, you can do so from here.
You can browse public results of FUMA (including example jobs) from Browse Public Results without registration or login.
Please post any questions, suggestions and bug reports on Google Forum: FUMA GWAS users.
Citation
When using SNP2GENE or GENE2FUNC modules, please cite the following:
K. Watanabe, E. Taskesen, A. van Bochoven and D. Posthuma. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8:1826. (2017).
links
https://www.nature.com/articles/s41467-017-01261-5
When using Cell Type module, please cite the following:
K. Watanabe, M. Umicevic Mirkov, C. de Leeuw, M. van den Heuvel and D. Posthuma. Genetic mapping of cell type specificity for complex traits. Nat. Commun. 10:3222. (2019).
https://www.nature.com/articles/s41467-019-11181-1
Depending on which results you are going to report, please also cite the original study of data sources/tools used in FUMA
(references are available at links or
tutorial for the cell type specificity analysis for scRNA-seq data).